vanddraabe is an R package to identify and analyze conserved waters within crystallographic protein structures and molecular dynamics simulation trajectories. The vanddraabe package expands on WatCH and PyWATER and returns statistical parameters for each water cluster, informative graphs, and a PyMOL session file to visually explore the conserved waters and protein. Hydrophilicity is the propensity of waters to congregate near specific protein atoms and is related to conserved waters. An informatics derived set of hydrophilicity values are provided based on a large, high-quality X-ray protein structure dataset.
vanddraabe is currently available on CRAN and GitHub.
Tutorials
The first tutorial (or vignettes in R) demonstrates the ability of vanddraabe to identify conserved waters within ten thrombin protein structures. This tutorial is based on the work of Sanschagrin and Kuhn (Protein Science, 1998, 7 (10), pp 2054-2064) and is part of the forthcoming vanddraabe article. (PDF | HTML)
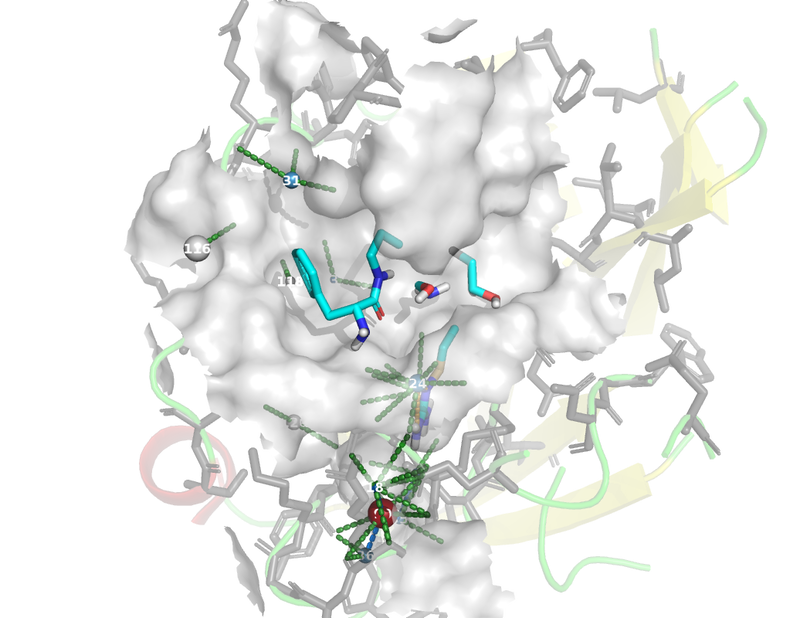
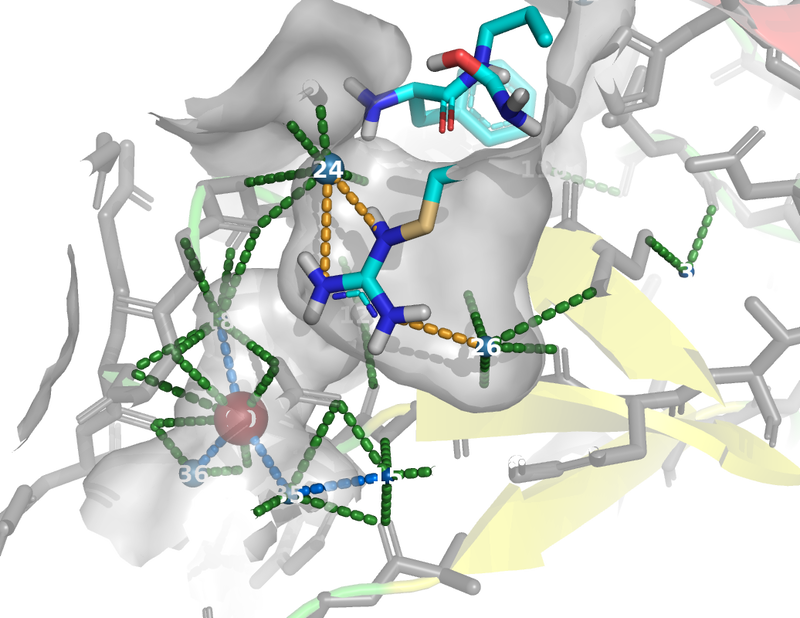
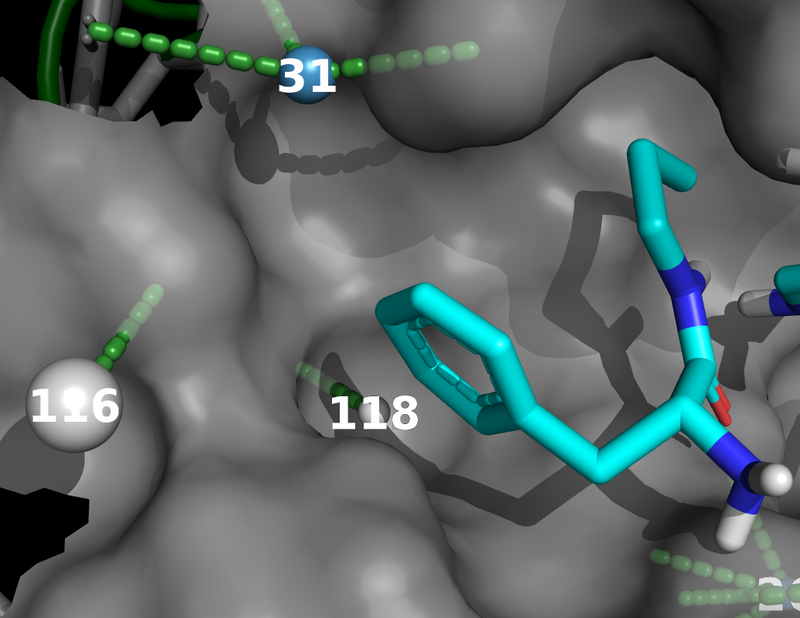
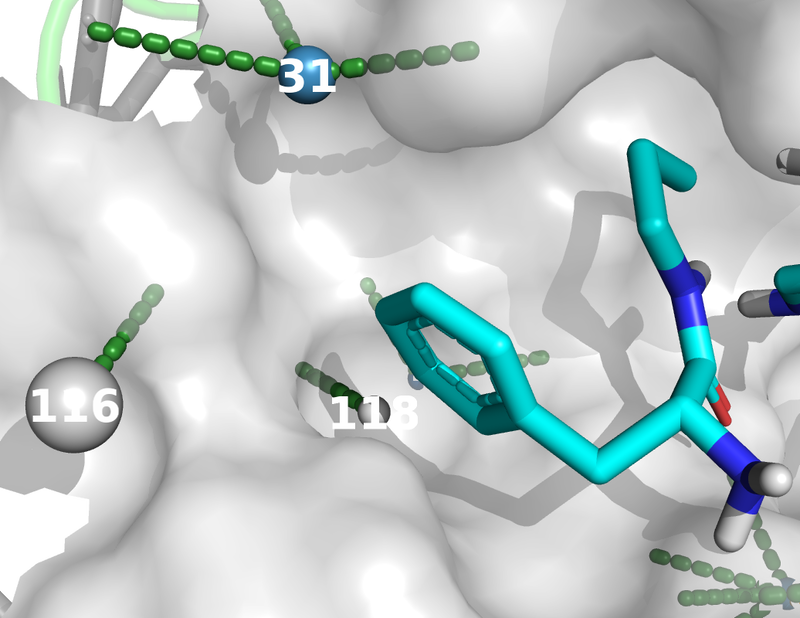
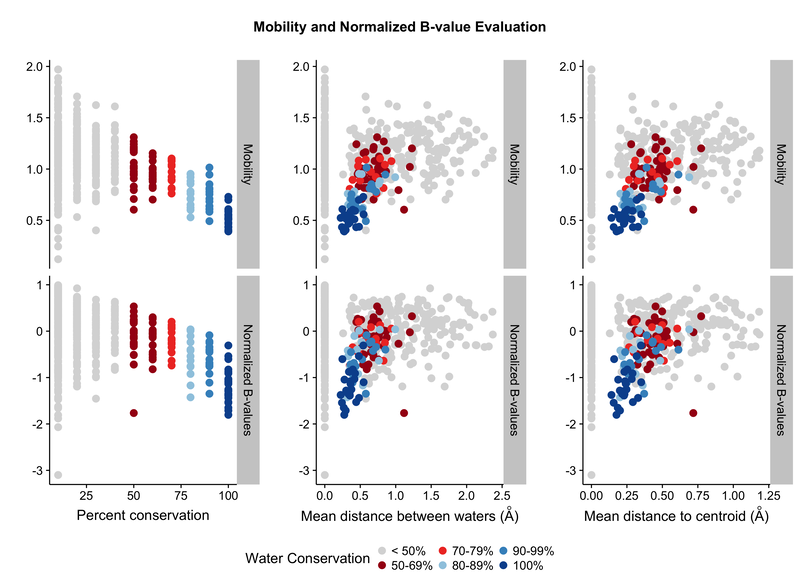
The images below were created from the tutorial.
vanddraabe is currently available on CRAN and GitHub.
Tutorials
The first tutorial (or vignettes in R) demonstrates the ability of vanddraabe to identify conserved waters within ten thrombin protein structures. This tutorial is based on the work of Sanschagrin and Kuhn (Protein Science, 1998, 7 (10), pp 2054-2064) and is part of the forthcoming vanddraabe article. (PDF | HTML)
The images below were created from the tutorial.